A Zebra Finch Basal Ganglia, and an Afternoon
A few months ago Rother, Januszewski, Jain, Fee, and Kornfeld released a preprint with an unusual claim: the first vertebrate basal ganglia connectome. Eight thousand five hundred neurons, twenty million synapses, a single zebra finch. I spent three years of my PhD analyzing connectomics datasets, and I know what it usually takes to get a new one into a form you can actually query. This one went online with a public REST API on day one.
I read the preprint in the morning. By that evening I had a clear finding.
This post is about what the paper changes, what I added in one afternoon, and why that time scale is starting to feel normal.
Songbirds learn to sing the way humans learn language
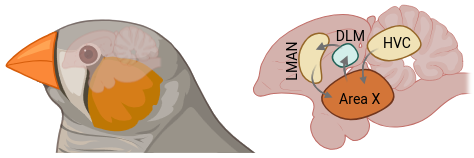
A juvenile male zebra finch hears his father sing, and over the next ninety days he learns to sing it back. He starts with babble, finds recognizable syllables, refines timing and pitch, and eventually produces a song so close to his tutor's that you can overlay the spectrograms. Then it locks in for life. This looks a lot like how a toddler learns to say spaghetti instead of pasghetti: a sensitive period, a template, auditory feedback, and a long arc of refinement. Songbirds are the best animal model we have for vocal learning, and zebra finches are the best songbird for it.
The circuit that does the learning is called Area X, and it sits in a bird-brain version of the basal ganglia — the same subcortical structures that Parkinson's disease takes apart, the same ones that learn to ride a bike or touch-type in humans. HVC feeds Area X a precise timing signal. LMAN feeds it exploratory noise. Dopamine from the midbrain tells it whether the last rendition was better or worse. The circuit gradually biases itself toward renditions that match the tutor. This is reinforcement learning, implemented in cells.
The textbook says two pathways, two cell types
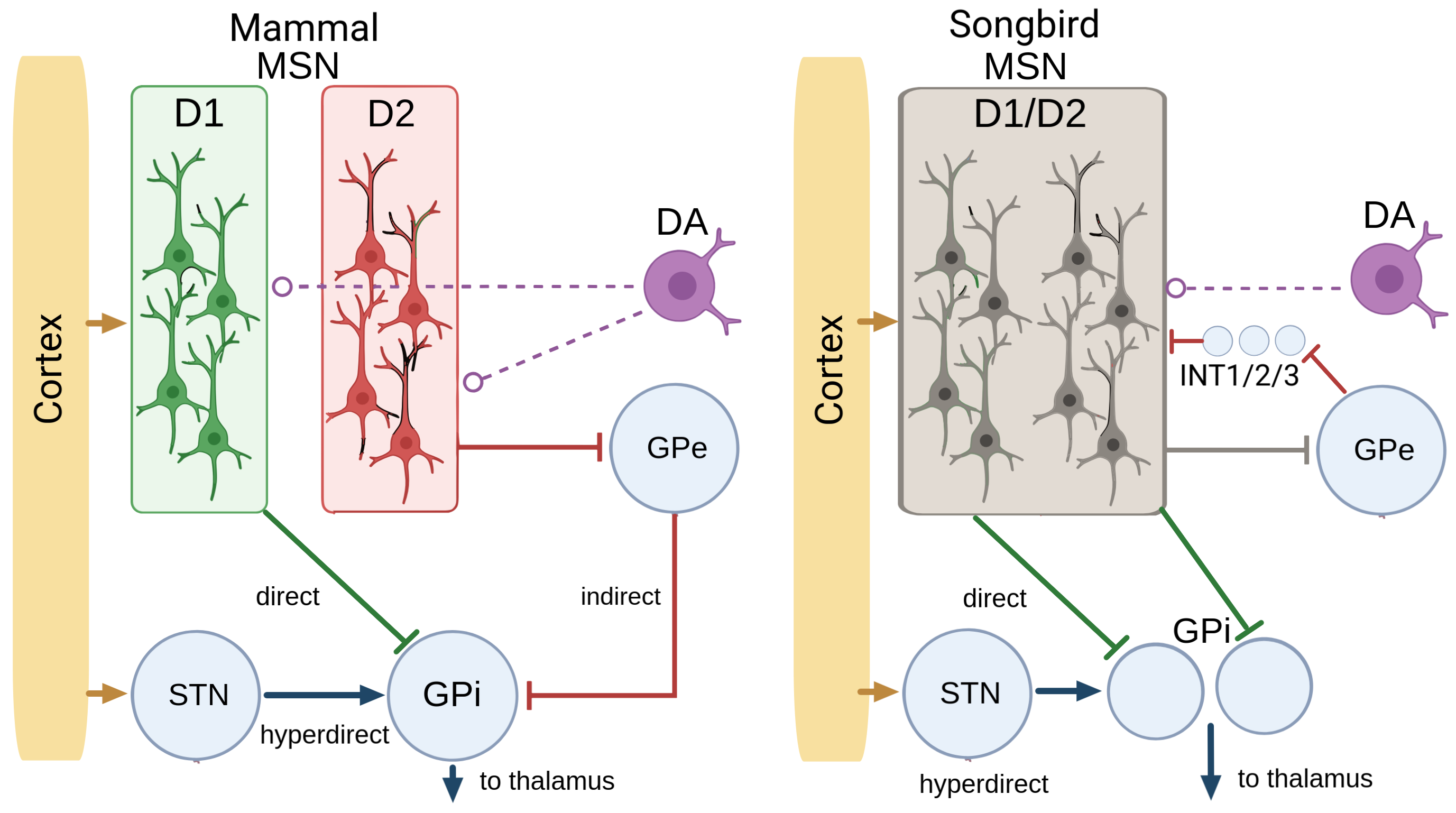
Every textbook model of the basal ganglia rests on a split. Two populations of neurons in the striatum, the input layer. One expresses the D1 dopamine receptor and drives a "go" pathway that promotes actions. The other expresses D2 and drives a "no-go" pathway that suppresses them. Dopamine tunes the balance.
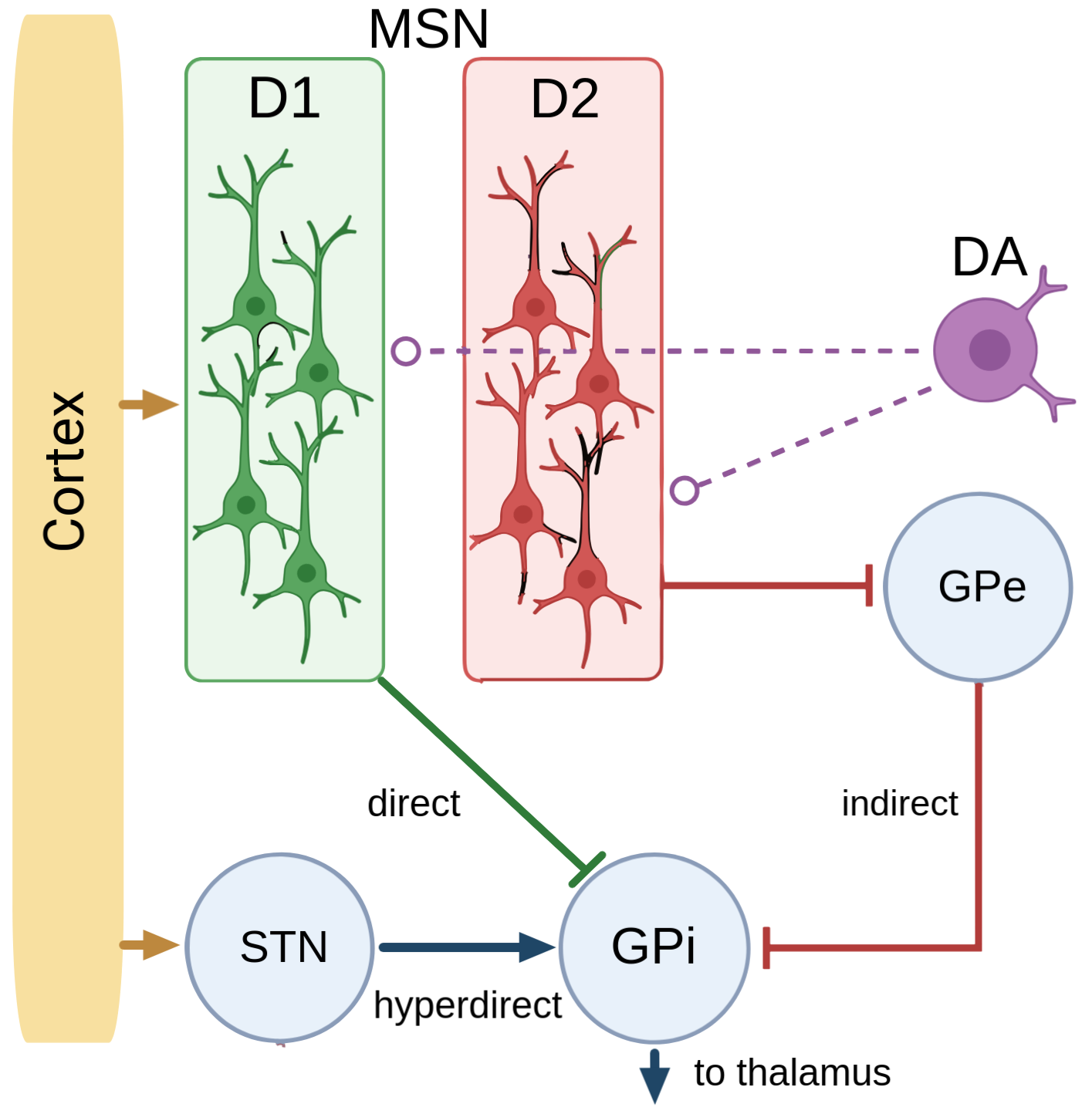
This model has organized forty years of work on movement, reward, and Parkinson's. It makes one strong assumption: the two pathways live in two anatomically distinct cell types.
In songbirds, that assumption had always been shaky. People had looked for D1 and D2 populations in Area X and come back with equivocal answers. Rother et al.'s connectome settled it. The D1/D2 distinction doesn't exist in Area X. MSNs are a single population that co-expresses both receptors. No two cell types, no direct-versus-indirect split at the cellular level.
So either songbirds don't do what the theory says they do, or the theory is wrong about which part is load-bearing, or the two-pathway logic lives somewhere else in the circuit. I wanted to know which.
One afternoon
The SyConn API went up with the preprint. Within two hours, I integrated the API into my connectomics MCP server and started querying. I pulled 800 MSNs and their output synapses onto the two pallidal populations (GPi and GPe) — the two downstream targets that correspond to the direct and indirect pathways in mammals. If the pathway distinction has moved off the cell-type axis, it should show up as structure in who targets what.
First I checked whether MSN output preferences split into two modes. A direct/indirect resurrection at the connectivity level would look like two peaks: some MSNs target mostly GPi, others mostly GPe. They don't. The distribution is a single right-skewed mode with a long tail toward the pure-direct end. Hartigan's dip test gave p-values above 0.05 across thresholds. The D1/D2 collapse holds at the output level too.
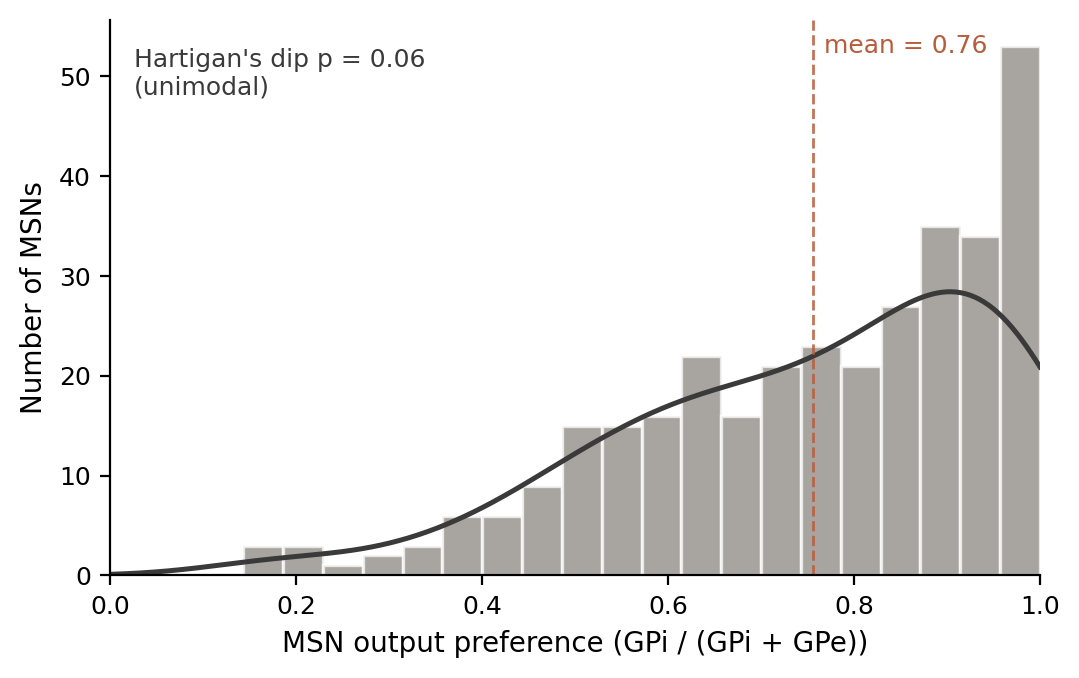
Then I tested whether the continuous variation — which is real and about five times what sampling noise would produce — tracks anything I could measure. HVC input fraction, LMAN input fraction, dopamine input, acetylcholine input, soma position, dendritic length. Nothing survived multiple-comparison correction. I then tested cell size using eight different size estimators, because the earlier hint of a size effect turned out to be synaptic density masquerading as size. Eight nulls.
At this point I thought I had a well-controlled negative result. The D1/D2 distinction is gone, there's no structural axis that replaces it, and the post I was going to write was going to be about how thoroughly the mammalian framework fails to transfer.
Then I flipped the question. I'd been asking what predicts which MSNs target which pallidal population. The other direction is: do the 47 GPi cells and 27 GPe cells receive from different mixtures of MSNs?
They do, and it isn't close. Individual GPi cells draw from MSN populations whose output preferences vary three to eight times more than you'd expect from uniform sampling. Two GPi cells sitting next to each other can be reading from completely different pools.
And the GPi cells themselves split into two groups. One group draws from MSNs across the full range of preferences, and its mean input profile matches GPe closely. The other group draws predominantly from MSNs at the direct-pathway end, and receives extra input from the subthalamic nucleus — which is the classical "hyperdirect" pathway input in mammals, carrying cortical signals straight to the output stage.
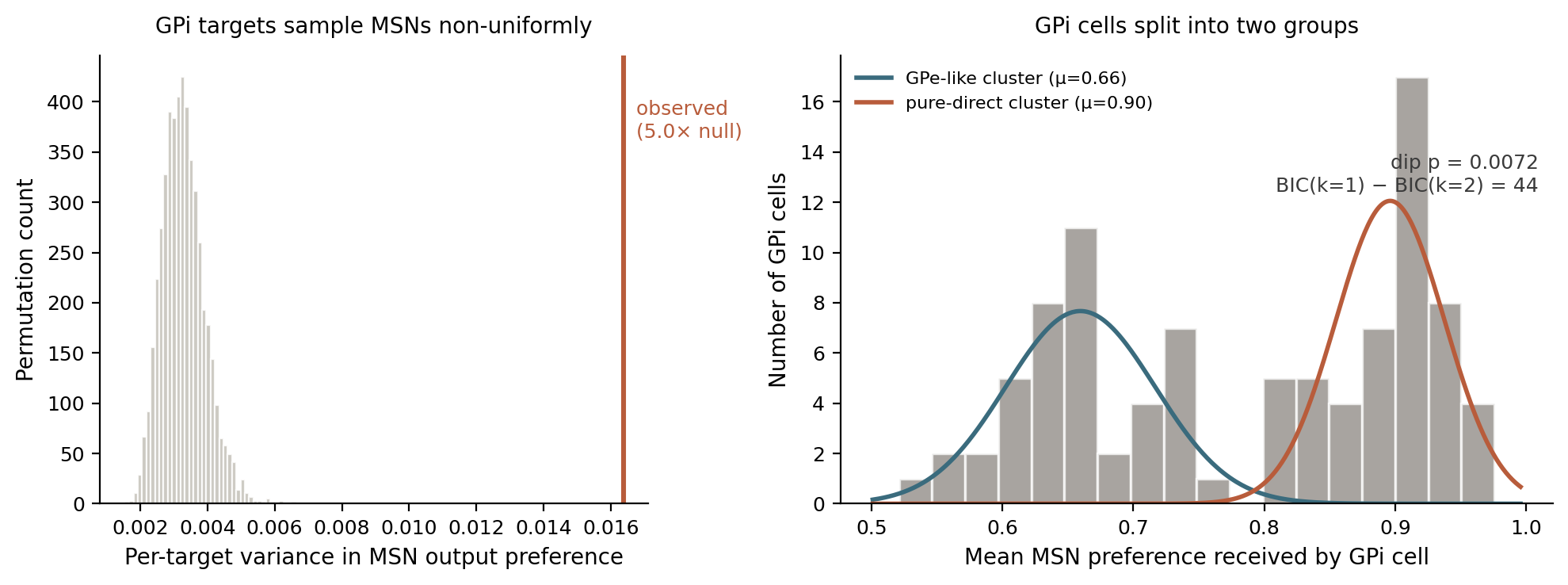
That second group is the direct+hyperdirect convergence motif from every mammalian basal ganglia textbook. In the mammalian textbook model, it's organized by cell type on the input side. In songbirds the input side is a single cell type, and the architecture gets rebuilt on the output side. Same computation, different parts.
The whole thing took about three hours end to end. Some of that was reading the preprint and integrating it with my MCP. The analysis that produced the headline finding was under an hour.
What I can say and what I can't say
I can say the GPi bimodality is robust. The dip test holds from n=25 to n=93 GPi cells, BIC favors a two-component mixture by 21 to 44 points at every threshold, and the separation between the modes is around five pooled standard deviations. The STN-input enrichment in the pure-direct cluster survives multiple-comparison correction. The lower GPi cluster and the GPe distribution overlap almost exactly in their mean MSN input profile, which suggests one GPi subpopulation is doing something GPe-like and the other is doing something distinctive to Area X.
I can't say from the public data which cluster is the one that projects out of Area X to drive thalamus. The authors have that classification internally, based on thick myelinated exit axons at the ultrastructural level. I tried to recover it from skeleton geometry — branch density, tortuosity, distance from arbor endpoints to the volume boundary — and the scale-invariant signal pointed the opposite direction from what the mammalian analogy would predict, but I can't rule out reconstruction-completeness confounds on the absolute-size features. Either answer is interesting.
The afternoon
I want to be honest about what this felt like. I didn't know I was going to find anything. I read the preprint while browsing bioRxiv. The three hours from "I should look at this" to "I think I've recovered the two-pathway logic on the output side" were not planned.
Five years ago this analysis would have taken months. The connectome wouldn't have existed. If it had, and was shared publically, you'd have written custom code to parse a terabyte of EM segmentations. If that had worked, you'd have spent weeks refining cell-type classifications. None of those steps are where the time goes anymore. Rother et al. built a public API, and I plugged that into my connectomics MCP to return structured data into my LLM's context window with just another couple hours. The cell types are pre-classified. The synapse detection is automated. The hard part is asking the right question and interpreting what comes back.
The Rother et al. dataset is small by flagship-connectome standards — fruit fly brains have ~140,000 neurons, mouse visual cortex projects target millions — but it's dense, well-curated, and released in a form you can query. Those three things together are what make afternoon-scale science possible.
References
- Rother, A., Januszewski, M., Jain, V., Fee, M.S., and Kornfeld, J. "The songbird basal ganglia connectome." _bioRxiv_2025.10.25.684569 (2025). doi:10.1101/2025.10.25.684569
- Fee, M.S., and Goldberg, J.H. "A hypothesis for basal ganglia-dependent reinforcement learning in the songbird." Neuroscience 198, 152–170 (2011). doi:10.1016/j.neuroscience.2011.09.069
- Person, A.L., Gale, S.D., Farries, M.A., and Perkel, D.J. "Organization of the songbird basal ganglia, including Area X." Journal of Comparative Neurology 508, 840–866 (2008). doi:10.1002/cne.21699
- Calabresi, P., Picconi, B., Tozzi, A., Ghiglieri, V., and Di Filippo, M. "Direct and indirect pathways of basal ganglia: a critical reappraisal." Nature Neuroscience 17, 1022–1030 (2014). doi:10.1038/nn.3743
- Nambu, A., Tokuno, H., and Takada, M. "Functional significance of the cortico-subthalamo-pallidal 'hyperdirect' pathway." Neuroscience Research 43, 111–117 (2002). doi:10.1016/S0168-0102(02)00027-5
- Schubert, P.J., Dorkenwald, S., Januszewski, M., Klimesch, J., Svara, F., Mancu, A., Ahmad, H., Fee, M.S., Jain, V., and Kornfeld, J. "SyConn2: dense synaptic connectivity inference for volume electron microscopy." Nature Methods 19, 1367–1370 (2022). doi:10.1038/s41592-022-01624-x