Demos from my Connectomics MCP
I spent three years of my PhD anaylyzing connectomics datasets: FlyWire, MICrONS, the hemibrain. Each one has its own API, its own naming conventions, its own way of making a small question take a whole day to answer. A simple question like "what are the inputs to this neuron?" requires reading docs, writing boilerplate, and reformatting results, all when you started by being curious about the biology hours ago.
I built an MCP server that lets Claude Code (CC) query and analyze all three datasets in the span of seconds. With it, I had the fly's locus of spatial memory, its heading-direction and path-integration circuits, mapped out in 6 minutes. I had a disinhibitory pathway in mouse V1 traced from an interneuron through its targets correlated to those targets' orientation tuning in under 9 minutes.
Here's my GitHub link if you want to try.
And here are those two explorations, if you're curious:
Spatial Memory in the Fly
My prompt into CC:
MeAsking about spatial memory
I want to better understand the spatial memory circuitry of the fruit fly. I've heard that it uses the connectivity of the ring neurons in the central complex to instantiate a 1-D ring attractor, and that motion signals it perceives like velocity and head direction help update its internal model of its position. I want to use the fly's hemibrain connectome to analyze that circuitry to understand the mechanism better. I don't know all the names of the relevant cells, unfortunately, so you'll have to dig that up yourself. Remember that this is still a field of open research, though it has been investigated for decades. So base your investigation on the connectome primarily, rather than textbook scientific descriptions.
And it's response:
ClaudeGetting started
I'll investigate the central complex ring attractor circuitry using the hemibrain connectome. Let me start by searching for the key cell types involved — EPG (compass/head direction), PEN (abgykar velocity), Delta7 (bump shaping), and ring neurons. Let me pull all of these in parallel.
CC made a few dozen tool calls over the next several minutes, tracing connectivity for each cell type and following the strongest connections downstream. Here's some of the comments it made along the way:
ClaudeFinding cells
All cell types are in the central complex. Let me now load the neuron tables and examine the actual connectivity between these types by picking representative neurons.
ClaudeTracing the ring
Excellent data. Now let me examine the critical topographic structure - how PEN neurons connect to EPG with a columnar offset (which enables the velocity integration). Let me also check what EPG projects to downstream.
ClaudeVerifying the columnar offset
The columnar structure is now very clear. Let me verify the criticall PEN-EPG offset wiring that drives bump rotation, by examining which PB glomeruli the connections occur in.
And then produced this summary:
ClaudeDrosophila Central Complex Ring Attractor: Hemibrain Connectome Analysis
## Drosophila Central Complex Ring Attractor: Hemibrain Connectome Analysis ### 1. The Compass — EPG Neurons (46 neurons) EPG neurons tile the **ellipsoid body (EB)** across 16 columnar positions: **L1–L8** and **R1–R8** in the protocerebral bridge (PB). A localized "bump" of activity in this population encodes the fly's internal estimate of head direction. Each column has 2–4 neurons (redundancy). All 46 neurons reside in the central complex (CX). ### 2. Angular Velocity Integration — PEN Neurons PEN_a neurons (20) and PEN_b neurons (22) both span columns L2-L9 and R2-R9 of the protocerebral bridge and integrate angular velocity signals. **The critical offset wiring:** Both PEN_a and PEN_b connect to EPG with a **-1 column offset** — a PEN neuron in column N excites EPG in column N-1. This is extremely consistent: - **PEN_a → EPG same-side offset: -1.03** (3,839 of 6,561 synapses at exactly -1; 1,859 at -2) - **PEN_b → EPG same-side offset: -1.07** (4,933 of 5,897 synapses at -1; 1,174 at -2) **EPG feeds back to PEN at offset ~+1** (EPG_col → PEN_col+1), completing a recurrent shift-and-amplify loop. When PEN neurons on one side are more active, the bump is pushed one step around the ring; the opposite side pushes it the other way. This is the core mechanism for **path integration of angular velocity**. ### 3. The Velocity Signal — GLNO Neurons (4 neurons) GLNO neurons (labeled `GLNO(LAL-NO1)`, 2 per hemisphere) are the **strongest external input to PEN neurons**: - → PEN_a: **4,199 synapses** - → PEN_b: **3,460 synapses** They connect the **lateral accessory lobe (LAL)** to the **nodulus (NO)** — a known pathway for relaying angular velocity signals derived from optic flow and mechanosensory inputs. Their own top inputs include PEN feedback (1,911 syn from PEN_a) and LAL interneurons (PS196, LAL139, LAL184), suggesting they are a key relay node for rotational velocity information. Other significant PEN inputs include: - **ExR6** (3,822 syn → PEN_a) and **ExR4** (3,398 syn → PEN_b) — extrinsic ring neurons likely providing modulatory/gain control - **LPsP** (2,871 syn → PEN_a), **IbSpsP** (2,565 → PEN_b, 1,701 → PEN_a) — lateral inputs possibly carrying translational velocity ### 4. Global Inhibition / Bump Shaping — Delta7 (42 neurons) Delta7 neurons are the **inhibitory backbone** that keeps the bump single and sharp. Key properties from the connectome: - **Massive recurrence:** 24,615 syn Delta7 → Delta7 - **Strong EPG drive:** 17,286 syn EPG → Delta7 (Delta7 reads the current bump) - **Inhibitory output to EPG:** 6,664 syn Delta7 → EPG - **Inhibition of PEN_b:** 9,800 syn Delta7 → PEN_b (but only 2,754 → PEN_a) **Antipodal column spanning:** Each Delta7 neuron innervates two PB columns that are roughly diametrically opposite on the ring (e.g., L2+R7, L3+R6, L4+R5, L5+R4, etc. — always summing to ~9). This creates the **surround inhibition** needed to suppress activity away from the bump, maintaining a single stable peak. This is the structural implementation of the "cosine-shaped" inhibition kernel predicted by ring attractor theory. ### 5. Visual Landmark Anchoring — Ring Neurons (ER types) Ring neurons (ER = ellipsoid body ring neurons) provide the **sensory anchoring** that corrects drift and ties the internal compass to external landmarks. The strongest ER inputs to EPG, by synapse count: - **ER4m** — 14,903 syn, visual feature input - **ER2_c** — 12,671 syn, visual feature input - **ER4d** — 10,010 syn, visual feature input - **ER3p_a** — 7,053 syn, visual feature input - **ER1_a** — 6,127 syn, visual + PEN_a input (989 syn) - **ER3w_b** — 5,875 syn, visual feature input - **EL** — 5,837 syn, ellipsoid body local Notably, some ring neuron subtypes show **target specificity** beyond EPG: - **ER1_a/ER1_b** → PEN_a preferentially (989 + 1,474 syn) - **ER6** → PEN_b preferentially (1,186 syn) - **ER4d** → PEN_b (786 syn) This suggests ring neurons don't just correct the compass — some subtypes may modulate the velocity integration gain directly. ### 6. Readout — EPG Downstream Targets EPG broadcasts the compass signal to diverse downstream circuits: - **EL** — 9,762 syn, ellipsoid body local processing - **ExR6** — 6,870 syn, extrinsic ring neuron (also feeds PEN) - **PEG** — 5,765 syn, same-column feedback to EPG (reinforcement) - **PFNd** — 2,653 syn, path integration / goal direction - **PFL2/PFL3** — 1,596/1,569 syn, steering commands (fan-shaped body → LAL → motor) - **PFNa** — 1,142 syn, navigation vector encoding The **PFL neurons** (protocerebral bridge → fan-shaped body → lateral accessory lobe) are particularly important — they read the compass and translate it into motor commands for steering.
ClaudeKey Mechanistic Insights from the Connectome
### Key Mechanistic Insights from the Connectome
1. **The offset is symmetric, not asymmetric by side.** Both L and R PEN neurons show the same -1 offset to EPG. The directional selectivity (clockwise vs counterclockwise rotation updating) likely comes from **differential activation** of left vs right PEN populations by GLNO, not from structural wiring asymmetry. Left GLNO drives left PEN, right GLNO drives right PEN, and both shift the bump in the same structural direction — but because L and R PB halves map to the EB ring in opposite orientations, the same -1 offset produces opposite rotational shifts on the ring.
2. **Delta7 provides structured, not uniform, inhibition.** The antipodal wiring (L_n + R_{9-n}) means each Delta7 neuron inhibits two diametrically opposite points on the ring, implementing a cosine-like inhibition kernel that's strongest opposite the bump and weakest at the bump itself.
3. **PEN_b is more strongly inhibited by Delta7** (9,800 syn) than PEN_a (2,754 syn), suggesting PEN_b may have a different gain regime or be gated more tightly.
4. **The circuit has multiple correction timescales**: fast angular velocity integration via PEN/GLNO, and slower visual landmark correction via the diverse ring neuron types (>20 subtypes), each presumably tuned to different visual features.
Six minutes and twenty-two seconds. That's all it took.
And the same question works with FlyWire. CC finds the same circuit (view in Neuroglancer) in about five minutes, though the experience is a little different — the hemibrain uses names from decades of Drosophila neuroscience (EPG, PEN, Delta7), while FlyWire has its own hierarchical taxonomy, and there's no universal mapping between them. Schlegel et al. 2023 built morphological correspondences across datasets using NBLAST scores, and that's in the pipeline for the MCP. For now, CC reads FlyWire's taxonomy and reasons about the correspondences itself. Imperfectly, but well enough to find the same circuit. FlyWire also has richer neurotransmitter annotations, which opens up questions the hemibrain can't answer, like how dopaminergic neurons modulate the goal vector computation.
The Functional Role of Disinhibition in Mouse Visual Cortex
The fly brain is a complete wiring diagram of a small nervous system. The MICrONS dataset reaches toward something bigger: a cubic millimeter of mouse visual cortex, reconstructed at synaptic resolution, with thousands of neurons that were also recorded in an awake mouse watching movies before the tissue was imaged. Structure and function, coregistered in the same cells. No other dataset can do this. The questions here are different from the fly. Instead of tracing uniquely identified neurons through a circuit, you're looking for connectivity motifs that repeat across the tissue. And because you have both wiring and visual responses, you can ask whether those motifs mean something.
I approached this one differently from the fly circuit. Instead of asking CC to characterize a known circuit, I wanted to see if it could connect two results from the MICrONS Nature suite to find something new: (1) the inhibitory motif groups from Schneider-Mizell et al., and (2) the coregistered functional imaging. The conversation revealed where expert scientific judgement still has a role to play.

MeAsking about disinhibition
Using the MICrONS dataset, find me an inhibitory neuron that Schneider-Mizell et al. identified as a disinhibitory specialist (one that preferentially targets other inhibitory neurons, specifically basket cells). Get its connectivity and tell me what it targets.
MeInhibiting basket cells to disinhibit pyramidal neurons
Pull the output connectivity of those basket cells and tell me: which pyramidal neurons do they inhibit, how many of those have coregistered functional data?
MeTuning of disinhibition (via basket cells)
Are the pyramidal neurons that are inhibited by those basket cells functionally similar to each other, or diverse? Compare the distribution of their tuning to a random sample of L2/3 pyramidal neurons in the same spatial neighborhood.
And after a grand total of 8 minutes, CC fetched the functional properties for thousands of neurons and wrote and ran some quick Python code to analyze their functional similarity.
ClaudeResults: Orientation Tuning of BC-Inhibited Pyramidal Neurons
## Results: Orientation Tuning of BC-Inhibited Pyramidal Neurons ### The short answer: functionally diverse, not clustered — but with subtle differences from controls. ### Orientation Preference - Both BC targets and neighborhood controls show a strong cardinal bias peaking at ~90° (vertical/horizontal orientations), which is typical of mouse V1. - The distributions are broadly similar, but BC targets (n=2,562) have a slightly tighter concentration around 72–108° (KS test p = 7×10⁻⁵). The circular variance confirms this: 0.69 for BC targets vs. 0.76 for controls — the BC targets are slightly less diverse in their orientation preferences. ### Selectivity Indices | Metric | BCs | Control | Signif. | |--------|-------------|-------------|-----------| | gOSI | 0.23 ± 0.13 | 0.22 ± 0.13 | p = 0.005 | | OSI | 0.42 ± 0.20 | 0.40 ± 0.20 | | | DSI | 0.23 ± 0.16 | 0.24 ± 0.17 | | BC-targeted pyramidal neurons are marginally more orientation-selective (higher gOSI/OSI) and slightly less direction-selective (lower DSI) than their neighbors (control, n=2,505, Mann-Whitney U-test). The effects are statistically significant given the large sample sizes but modest in magnitude (~5% relative difference). ### Per-Basket-Cell Diversity The bottom-right panel shows that every individual basket cell targets a diverse pool of orientations (circular variance 0.65–0.80 across all 5 BCs) — none of them selectively inhibits a single orientation column. ### Interpretation This is consistent with the known physiology of PV+ basket cells in mouse V1: they provide broad, non-selective perisomatic inhibition (the "blanket of inhibition" model). Each BC contacts thousands of local pyramidals spanning the full range of orientation preferences. The slight enrichment in orientation selectivity among BC targets may reflect that basket cells preferentially contact neurons with stronger sensory drive — which would make them better targets for gain control — but the effect is subtle. These BCs are not carving out orientation-specific subnetworks; they are performing divisive normalization across the local population.
So CC found that basket cells disinhibited by an ITC target a functionally diverse pool of L2/3 pyramidals. That's standard; PV+ basket cells provide broad perisomatic inhibition. But how about the Martinotti cells? 3 more minutes of CC and:
ClaudeMartinotti Cell Pathway: Key Findings
## Martinotti Cell Pathway: Key Findings ### MC vs BC: Strikingly similar — but with a critical structural difference The MC and BC pathways are functionally indistinguishable (KS tests: pref_ori p=0.88, gOSI p=0.99, DSI p=0.43). Both populations show: - Circular mean orientation ~82°, broad cardinal bias - Mean gOSI ~0.23, mean DSI ~0.22–0.23 - High circular variance (0.68–0.69) — diverse, not clustered
ClaudeTuning of shared (MC+BC) disinhibition
### The massive overlap is the real story 2,293 out of 3,226 MC-targeted L2/3 neurons (71%) are also targeted by the BCs. These two inhibitory pathways converge onto a largely shared pyramidal population, Shared (MC+BC). Breaking the targets into exclusive vs shared groups reveals a gradient: | Group | n | gOSI | CirVar | DSI | |---------|-------|-------|--------|-------| | Shared | 1,630 | 0.235 | 0.660 | 0.224 | | BC-only | 932 | 0.226 | 0.754 | 0.237 | | MC-only | 610 | 0.216 | 0.725 | 0.216 | | Control | 1,970 | 0.219 | 0.743 | 0.247 | The shared targets — neurons receiving convergent MC and BC inhibition — are the most orientation-selective (highest gOSI) and the most coherently tuned (lowest circular variance, CirVar). MC-only and BC-only targets look more like the general population. ### Interpretation Both the Martinotti and basket cell arms of this disinhibitory circuit target a largely overlapping, functionally diverse pool of L2/3 pyramidals — but the neurons that sit at the intersection of both inhibitory streams are slightly but consistently more tuned. This is consistent with a model where the most visually driven neurons in the local population receive the strongest inhibitory regulation from both dendritic (MC) and perisomatic (BC) compartments. The ITC disinhibitor, by releasing both brakes simultaneously, would preferentially boost these well-tuned neurons — a form of feature-selective gain modulation emerging from a structurally non-selective circuit.
That caught me off guard: Claude flagging that the neurons at the intersection of both the Martinotti and basket cell pathways were more orientation-coherent and more selective than the rest. That looked like a real circuit-specific signal: convergent dendritic and perisomatic inhibition landing on a functionally distinct subpopulation.
But let's get precise about the controls. The pyramidal neurons suppressed by both pathways exist in an even smaller spatial volume, and since the primary visual cortex is retinotopic, these neurons are even more likely to respond to similar orientations.
MeQuestioning the control
Can you explain the control you used here? Is there a need for a separate control for the Shared group as compared to the control for the BC-only or MC-only groups?
ClaudeThe control
## The control The control is a random sample of L2/3 pyramidal neurons from the same spatial neighborhood that are not targeted by any of the 5 BCs or 5 MCs in this analysis. "Same spatial neighborhood" is defined as neurons whose soma positions fall within ±2 standard deviations of the BC/MC target population's centroid in the x and z dimensions (the tangential cortical axes), without filtering on y (depth axis, which would conflate laminar identity with spatial proximity). This controls for: - Laminar identity (all are 23P) - Retinotopic position (same cortical patch, so similar visual field coverage) - Imaging coverage bias (same region is likely covered by the same 2-photon sessions)
ClaudeDoes the Shared group need a separate control?
## Does the Shared group need a separate control? Yes, arguably it does, and the current analysis has a confound. The issue is: Neurons targeted by more interneurons are, by definition, more synaptically connected. A neuron that receives convergent BC+MC input likely has a larger dendritic arbor, more total synapses, or sits in a denser part of the neuropil. These are neurons that are more "visible" to the connectome — and being more connected could correlate with being more visually driven (higher firing rates → more orientation selectivity) for purely structural reasons, not because of any specific circuit logic. A proper control for the Shared group would be L2/3 pyramidals that are: 1. In the same spatial neighborhood 2. Matched in total inhibitory synapse count (or total postsynaptic count) — i.e., they receive a comparable amount of inhibition, just not from these specific MCs and BCs 3. Not targeted by any of the 10 interneurons in this analysis Without that, the gOSI difference in the Shared group (0.235 vs 0.219 in control) could simply reflect a connectivity-selectivity correlation: neurons with more synaptic partners tend to be more orientation-selective because they're larger, healthier, or better-proofread neurons in the EM volume. After matching controls on neuron volume (a proxy for total connectivity/arbor size), the picture changes substantially from the original analysis:
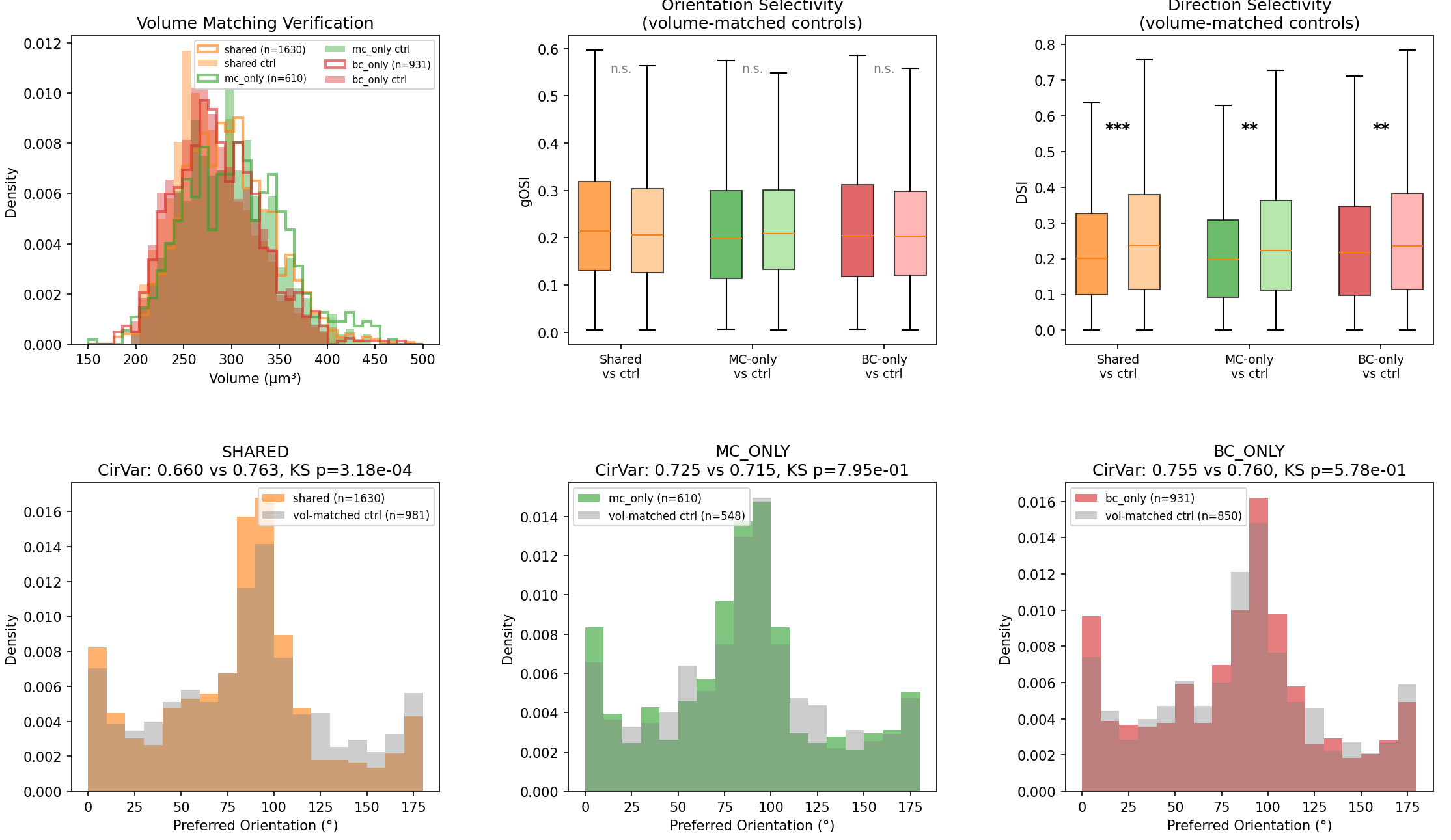
ClaudeOrientation Selectivity (gOSI): No longer significant
### Orientation Selectivity (gOSI): No longer significant The gOSI differences we saw earlier do not survive volume matching. The Shared group's elevated selectivity (which looked interesting before) shrinks to a non-significant trend (p = 0.10, d = +0.08). This suggests the original effect was at least partly driven by the connectivity-selectivity confound we anticipated.
ClaudeDirection Selectivity (DSI): Robust and real
### Direction Selectivity (DSI): Robust and real The one signal that does survive is in direction selectivity — all three target groups show lower DSI than their volume-matched controls (Shared: p = 1.3×10⁻⁴, MC-only: p = 4.2×10⁻³, BC-only: p = 8.8×10⁻³). These interneurons preferentially target pyramidal neurons that are less direction-selective than their neighbors of the same size.
ClaudeOrientation preference (CirVar): Shared group shows tighter tuning
### Orientation preference (CirVar): Only the Shared group shows tighter tuning The tighter clustering of preferred orientation survives in the Shared group (CirVar 0.660 vs 0.764, KS p = 3.2×10⁻⁴) but not in MC-only (p = 0.80) or BC-only (p = 0.58). This means the orientation coherence is specific to the neurons receiving convergent MC+BC inhibition — not a generic property of being targeted by any one interneuron.
ClaudeBottom line
### Bottom line The volume-matched controls reveal that: 1. The gOSI enrichment was largely a connectivity confound — more-connected neurons tend to be more orientation-selective, and that explained most of the effect. 2. The DSI depression is real — these interneurons preferentially target orientation-tuned but not direction-tuned pyramidals, regardless of neuron size. 3. The orientation coherence in the Shared group is the one genuinely circuit-specific signal — neurons at the intersection of dendritic (MC) and perisomatic (BC) inhibition share more similar orientation preferences than you'd expect by chance, even after controlling for volume.
So the orientation selectivity enrichment didn't survive volume matching — more-connected neurons tend to be more selective, and that explained most of the effect. But two signals did survive: the shared MC+BC targets still had tighter orientation coherence (p = 3.2×10⁻⁴), and all three target groups showed lower direction selectivity than their size-matched neighbors. These interneurons preferentially target neurons that are orientation-tuned but not direction-tuned, regardless of how connected those neurons are.
Conclusion
Is this a discovery? Not from a single ITC neuron. A proper analysis would need to trace this across many ITC circuits and account for each neuron's proofreading quality — both queryable through the MCP, both natural next steps. But going from a structural motif to a functional hypothesis to a controlled statistical test in a single CC session, with the MCP supplying data at every step — that's the workflow I wanted to exist three years ago.
One thing worth saying explicitly: I checked CC's work. The data retrieval is transparent (every tool call and what came back is logged right there in front of you) so auditing the inputs is straightforward. And the reasoning over that data is checkable against the tables CC itself produced. The analysis code is where I'd be most cautious. CC writes Python, runs stats, and generates figures, and bugs in analysis code can be silent, producing results that look right but aren't. I verified the results in these demos, and the push-and-pull over controls in the MICrONS section is a real example of what that looks like in practice. But this is no different from reviewing a colleague's analysis or your own code from last month. None of this is unique to AI. Mistakes happen in every analysis pipeline. The relevant question is whether the workflow makes errors easy to find.
The MCP is at GitHub link and lets Claude Code query the three major connectomics datasets (FlyWire, the hemibrain via neuPrint, and MICrONS via CAVE, including the functional imaging data). I'm still building it — bulk connectivity downloads, cross-dataset cell type mapping via NBLAST scores, and better coverage of the MICrONS annotation tables are next. I've used the H01 dataset before too, but it's even less accessible. If you work with connectomics, try it out and tell me what's missing.
References
Dorkenwald, S. et al. "Neuronal wiring diagram of an adult brain." Nature 634, 124–138 (2024). doi:10.1038/s41586-024-07558-y
Scheffer, L.K. et al. "A connectome and analysis of the adult Drosophila central brain." eLife 9, e57443 (2020). doi:10.7554/eLife.57443
MICrONS Consortium et al. "Functional connectomics spanning multiple areas of mouse visual cortex." Nature 640, 435–447 (2025). doi:10.1038/s41586-025-08790-w
Schneider-Mizell, C.M. et al. "Inhibitory specificity from a connectomic census of mouse visual cortex." Nature 640, 448–458 (2025). doi:10.1038/s41586-024-07780-8
Schlegel, P. et al. "Whole-brain annotation and multi-connectome cell typing of Drosophila." Nature 634, 139–152 (2024). doi:10.1038/s41586-024-07686-5